8 Describing a profile with python and geni-lib
See the geni-lib manual
geni-lib is a tool that allows users to generate RSpec files from Python code. CloudLab offers the ability to use geni-lib scripts as the definition of a profile, rather then the more primitive RSpec format. When you supply a geni-lib script on the Create Profile page, your script is uploaded to the server so that it can be executed in the geni-lib environment. This allows the script to be verified for correctness, and also produces the equivalent RSpec representation that you can view if you so desire.
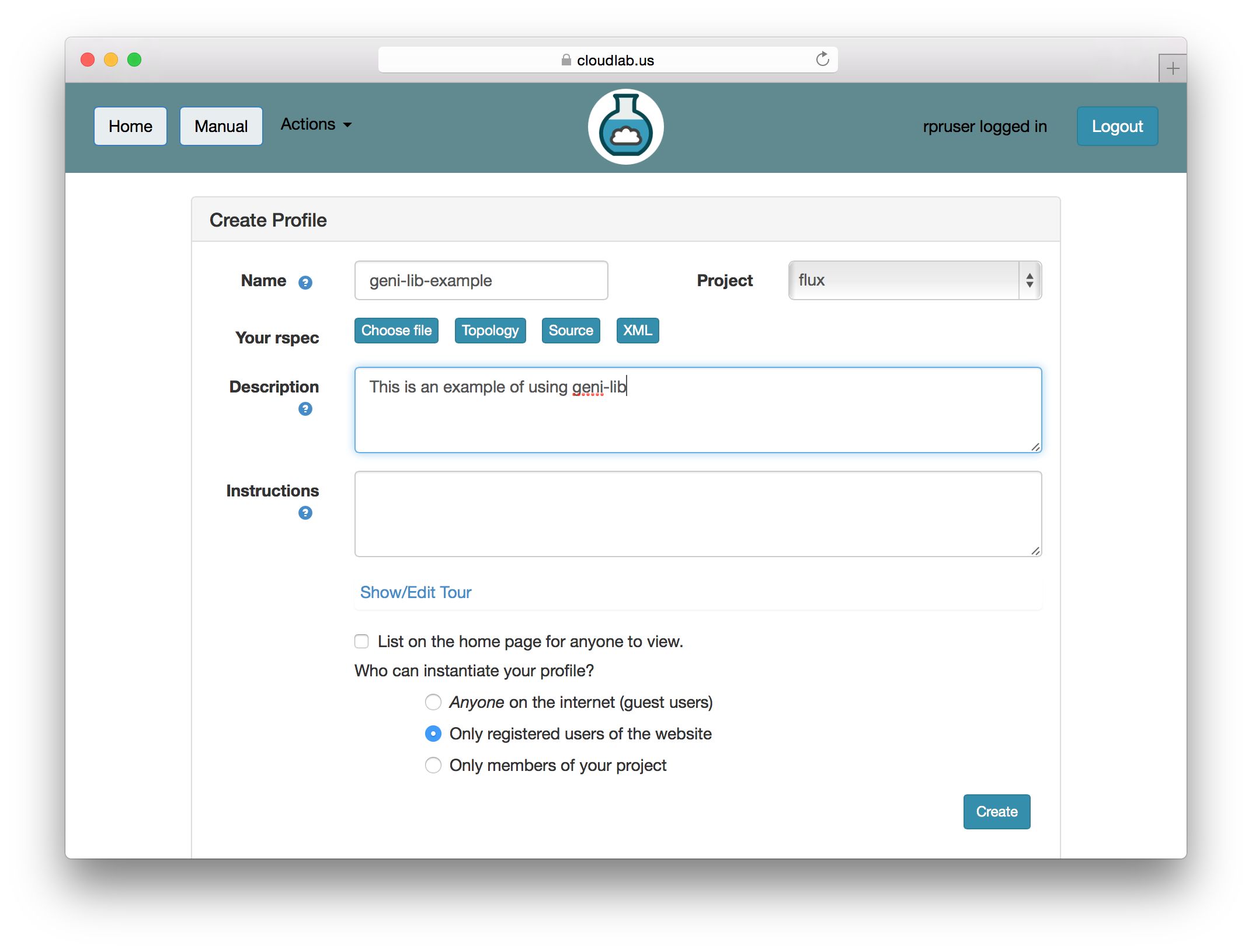
When you provide a geni-lib script, you will see a slightly different set of buttons on the Create Profile page; next to the “Source” button there is an “XML” button that will pop up the RSpec XML for you to look at. The XML is read-only; if you want to change the profile, you will need to change the python source code that is displayed when you click on the “Source” button. Each time you change the python source code, the script is uploaded to the server and processed. Be sure to save your changes if you are updating an existing profile.
The following examples demonstrate basic geni-lib usage. More information about geni-lib and additional examples, can be found in the geni-lib repository. Its full documentation is online as part of this manual.
8.1 A single XEN VM node
"""An example of constructing a profile with a single Xen VM. Instructions: Wait for the profile instance to start, and then log in to the VM via the ssh port specified below. (Note that in this case, you will need to access the VM through a high port on the physical host, since we have not requested a public IP address for the VM itself.) """ # Import the Portal object. import geni.portal as portal import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Add a XenVM (named "node") to the request node = request.XenVM("node") # Write the request in RSpec format portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of constructing a profile with a single Xen VM. Instructions: Wait for the profile instance to start, and then log in to the VM via the ssh port specified below. (Note that in this case, you will need to access the VM through a high port on the physical host, since we have not requested a public IP address for the VM itself.) """ # Import the Portal object. import geni.portal as portal import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Add a XenVM (named "node") to the request node = request.XenVM("node") # Write the request in RSpec format portal.context.printRequestRSpec()
This example demonstrates the two most important objects: the portal context (accessed through the portal.context object in the geni.portal module), and the request RSpec created by calling makeRequestRSpec() on it. These fundamental objects are central to essentially all CloudLab geni-lib profiles.
Another way to create a Request RSpec object is to call its constructuor, geni.rspec.pg.Request directly. We ask the Context to create it for us so it it is "bound" to the context and does not need to be explicitly passed to other functions on the context
Once the request object has been created, resources may be added to it by calling methods on it like RawPC() or rspec.pg.LAN. In this example, just a single node (created with the XenVM() constructor, asking for a single VM identified by the name "node") is requested.
Most functions called on Request objects are not directly members of that class. Rather, they are loaded as "extensions" by modules such as geni.rspec.emulab.
The final action the geni-lib script performs is to generate the XML representation of the request RSpec, with the printRequestRSpec() call on the last line. This has the effect of communicating the description of all the resources requested by the profile back to CloudLab.
You will also notice that the profile begins with a string literal (to be precise, it is a Python docstring). The initial text will also be used as the profile description; the text following the Instructions: line will be used as the corresponding instructions. This documentation is so important that adding the description to the profile is mandatory. (Using a docstring like this is not the only way to produce the description and instructions, although it is the most convenient.)
This simple example has now demonstrated all the important elements of a geni-lib profile. The portal context and request RSpec objects, the final printRequestRSpec() call, and the docstring description and instructions are “boilerplate” constructions, and you will probably include similar or identical versions of them in every geni-lib profile you create unless you are doing something quite unusual.
8.2 A single physical host
"""An example of constructing a profile with a single raw PC. Instructions: Wait for the profile instance to start, and then log in to the host via the ssh port specified below. """ import geni.portal as portal import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Create a raw PC node = request.RawPC("node") # Print the RSpec to the enclosing page. portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of constructing a profile with a single raw PC. Instructions: Wait for the profile instance to start, and then log in to the host via the ssh port specified below. """ import geni.portal as portal import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Create a raw PC node = request.RawPC("node") # Print the RSpec to the enclosing page. portal.context.printRequestRSpec()
As mentioned above, most of these simple examples consist of boilerplate geni-lib fragments, and indeed the portal context and request RSpec operations are unchanged from the previous script. The big difference, though (other than the updated documentation) is that in this case the RawPC() method is invoked on the Request object instead of XenVM(). As you might expect, the new profile will request a physical host instead of a virtual one. (A side effect of using a real machine is that it automatically comes with a unique public IP address, where the VM used in the earlier example did not. Profiles can request public IP addresses for VMs too, though it does not happen by default.)
8.3 Two XenVM nodes with a link between them
"""An example of constructing a profile with two VMs connected by a LAN. Instructions: Wait for the profile instance to start, and then log in to either VM via the ssh ports specified below. """ import geni.portal as portal import geni.rspec.pg as rspec request = portal.context.makeRequestRSpec() # Create two XenVM nodes. node1 = request.XenVM("node1") node2 = request.XenVM("node2") # Create a link between them link1 = request.Link(members = [node1,node2]) portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of constructing a profile with two VMs connected by a LAN. Instructions: Wait for the profile instance to start, and then log in to either VM via the ssh ports specified below. """ import geni.portal as portal import geni.rspec.pg as rspec request = portal.context.makeRequestRSpec() # Create two XenVM nodes. node1 = request.XenVM("node1") node2 = request.XenVM("node2") # Create a link between them link1 = request.Link(members = [node1,node2]) portal.context.printRequestRSpec()
This example demonstrates two important geni-lib concepts: first, adding more than a single node to the request (which is a relatively straightforward matter of calling more than one node object constructor, being careful to use a different name each time). It also shows how to add links between nodes. It is possible to construct links and LANs in a more complicated manner (such as explicitly creating Interface objects to control interfaces), but the simplest case is to supply the member nodes at the time the link is created.
8.4 Two ARM64 servers in a LAN
"""An example of constructing a profile with two ARM64 nodes connected by a LAN. Instructions: Wait for the profile instance to start, and then log in to either host via the ssh ports specified below. """ import geni.portal as portal import geni.rspec.pg as rspec request = portal.context.makeRequestRSpec() # Create two raw "PC" nodes node1 = request.RawPC("node1") node2 = request.RawPC("node2") # Set each of the two to specifically request "m400" nodes, which in CloudLab, are ARM node1.hardware_type = "m400" node2.hardware_type = "m400" # Create a link between them link1 = request.Link(members = [node1, node2]) portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of constructing a profile with two ARM64 nodes connected by a LAN. Instructions: Wait for the profile instance to start, and then log in to either host via the ssh ports specified below. """ import geni.portal as portal import geni.rspec.pg as rspec request = portal.context.makeRequestRSpec() # Create two raw "PC" nodes node1 = request.RawPC("node1") node2 = request.RawPC("node2") # Set each of the two to specifically request "m400" nodes, which in CloudLab, are ARM node1.hardware_type = "m400" node2.hardware_type = "m400" # Create a link between them link1 = request.Link(members = [node1, node2]) portal.context.printRequestRSpec()
We now come to demonstrate requesting particular properties of nodes—
8.5 A VM with a custom size
"""An example of constructing a profile with a single Xen VM. Instructions: Wait for the profile instance to start, and then log in to the VM via the ssh port specified below. (Note that in this case, you will need to access the VM through a high port on the physical host, since we have not requested a public IP address for the VM itself.) """ import geni.portal as portal import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Create a XenVM node = request.XenVM("node") # Ask for two cores node.cores = 2 # Ask for 2GB of ram node.ram = 2048 # Add an extra 8GB of space on the primary disk. # NOTE: Use fdisk, the extra space is in the 4th DOS partition, # you will need to create a filesystem and mount it. node.disk = 8 # Alternate method; request an ephemeral blockstore mounted at /mydata. # NOTE: Comment out the above line (node.disk) if you do it this way. #bs = node.Blockstore("bs", "/mydata") #bs.size = "8GB" #bs.placement = "nonsysvol" # Print the RSpec to the enclosing page. portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of constructing a profile with a single Xen VM. Instructions: Wait for the profile instance to start, and then log in to the VM via the ssh port specified below. (Note that in this case, you will need to access the VM through a high port on the physical host, since we have not requested a public IP address for the VM itself.) """ import geni.portal as portal import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Create a XenVM node = request.XenVM("node") # Ask for two cores node.cores = 2 # Ask for 2GB of ram node.ram = 2048 # Add an extra 8GB of space on the primary disk. # NOTE: Use fdisk, the extra space is in the 4th DOS partition, # you will need to create a filesystem and mount it. node.disk = 8 # Alternate method; request an ephemeral blockstore mounted at /mydata. # NOTE: Comment out the above line (node.disk) if you do it this way. #bs = node.Blockstore("bs", "/mydata") #bs.size = "8GB" #bs.placement = "nonsysvol" # Print the RSpec to the enclosing page. portal.context.printRequestRSpec()
The earlier examples requesting VMs used the default number of cores, quantity of RAM, and disk size. It’s also possible to customize these value, as this example does by setting the cores, ram, and disk properties of the XenVM class (which is a subclass of rspec.pg.Node.)
8.6 Set a specific IP address on each node
"""An example of constructing a profile with node IP addresses specified manually. Instructions: Wait for the profile instance to start, and then log in to either VM via the ssh ports specified below. (Note that even though the EXPERIMENTAL data plane interfaces will use the addresses given in the profile, you will still connect over the control plane interfaces using addresses given by the testbed. The data plane addresses are for intra-experiment communication only.) """ import geni.portal as portal import geni.rspec.pg as rspec request = portal.context.makeRequestRSpec() node1 = request.XenVM("node1") iface1 = node1.addInterface("if1") # Specify the component id and the IPv4 address iface1.component_id = "eth1" iface1.addAddress(rspec.IPv4Address("192.168.1.1", "255.255.255.0")) node2 = request.XenVM("node2") iface2 = node2.addInterface("if2") # Specify the component id and the IPv4 address iface2.component_id = "eth2" iface2.addAddress(rspec.IPv4Address("192.168.1.2", "255.255.255.0")) link = request.LAN("lan") link.addInterface(iface1) link.addInterface(iface2) portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of constructing a profile with node IP addresses specified manually. Instructions: Wait for the profile instance to start, and then log in to either VM via the ssh ports specified below. (Note that even though the EXPERIMENTAL data plane interfaces will use the addresses given in the profile, you will still connect over the control plane interfaces using addresses given by the testbed. The data plane addresses are for intra-experiment communication only.) """ import geni.portal as portal import geni.rspec.pg as rspec request = portal.context.makeRequestRSpec() node1 = request.XenVM("node1") iface1 = node1.addInterface("if1") # Specify the component id and the IPv4 address iface1.component_id = "eth1" iface1.addAddress(rspec.IPv4Address("192.168.1.1", "255.255.255.0")) node2 = request.XenVM("node2") iface2 = node2.addInterface("if2") # Specify the component id and the IPv4 address iface2.component_id = "eth2" iface2.addAddress(rspec.IPv4Address("192.168.1.2", "255.255.255.0")) link = request.LAN("lan") link.addInterface(iface1) link.addInterface(iface2) portal.context.printRequestRSpec()
This code sample assigns specific IP addresses to interfaces on the nodes it requests.
Some of the available qualifiers on requested nodes are specified by manipulating attributes within the node (or interface) object directly. The hardware_type in the previous example is one such case, as is the component_id here. (Note that the component_id in this example is applied to an interface, although it is also possible to specify component_ids on nodes, too, to request a particular physical host.)
Other modifications to requests require dedicated methods. For instance, see the addAddress() calls made on each of the two interfaces above. In each case, an IPv4Address object is obtained from the appropriate constructor (the parameters are the address and the netmask, respectively), and then added to the corresponding interface.
8.7 Specify an operating system and set install and execute scripts
"""An example of constructing a profile with install and execute services. Instructions: Wait for the profile instance to start, then click on the node in the topology and choose the `shell` menu item. The install and execute services are handled automatically during profile instantiation, with no manual intervention required. """ # Import the Portal object. import geni.portal as portal # Import the ProtoGENI library. import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Add a raw PC to the request. node = request.RawPC("node") # Request that a specific image be installed on this node node.disk_image = "urn:publicid:IDN+emulab.net+image+emulab-ops//UBUNTU20-64-STD"; # Install and execute scripts on the node. THIS TAR FILE DOES NOT ACTUALLY EXIST! node.addService(rspec.Install(url="http://example.org/sample.tar.gz", path="/local")) node.addService(rspec.Execute(shell="bash", command="/local/example.sh")) portal.context.printRequestRSpec()Open this profile on CloudLab"""An example of constructing a profile with install and execute services. Instructions: Wait for the profile instance to start, then click on the node in the topology and choose the `shell` menu item. The install and execute services are handled automatically during profile instantiation, with no manual intervention required. """ # Import the Portal object. import geni.portal as portal # Import the ProtoGENI library. import geni.rspec.pg as rspec # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Add a raw PC to the request. node = request.RawPC("node") # Request that a specific image be installed on this node node.disk_image = "urn:publicid:IDN+emulab.net+image+emulab-ops//UBUNTU20-64-STD"; # Install and execute scripts on the node. THIS TAR FILE DOES NOT ACTUALLY EXIST! node.addService(rspec.Install(url="http://example.org/sample.tar.gz", path="/local")) node.addService(rspec.Execute(shell="bash", command="/local/example.sh")) portal.context.printRequestRSpec()
This example demonstrates how to request services for a node, where CloudLab will automate some task as part of the profile instance setup procedure. In this case, two services are described (an install and an execute). This is a very common pair of services to request together: the Install object describes a service which retrieves a tarball from the location given in the url parameter, and installs it into the local filesystem as specified by path. (The installation occurs during node setup, upon the first boot after the disk image has been loaded.) The second service, described by the Execute object, invokes a shell process to run the given command. In this example (as is common), the command refers directly to a file saved by the immediately preceding Install service. This behaviour works, because CloudLab guarantees that all Install services complete before any Execute services are started. The command executes every time the node boots, so you can use it start daemons, etc. that are necessary for your experiment.
8.8 Profiles with user-specified parameters
"""An example of using parameters to construct a profile with a variable number of nodes. Instructions: Wait for the profile instance to start, and then log in to one or more of the VMs via the ssh port(s) specified below. """ # Import the Portal object. import geni.portal as portal # Import the ProtoGENI library. import geni.rspec.pg as rspec # Describe the parameter(s) this profile script can accept. portal.context.defineParameter( "n", "Number of VMs", portal.ParameterType.INTEGER, 1 ) # Retrieve the values the user specifies during instantiation. params = portal.context.bindParameters() # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Check parameter validity. if params.n < 1 or params.n > 8: portal.context.reportError( portal.ParameterError( "You must choose at least 1 and no more than 8 VMs.", ["n"] ) ) # Abort execution if there are any errors, and report them. portal.context.verifyParameters() for i in range( params.n ): # Create a XenVM and add it to the RSpec. node = request.XenVM( "node" + str( i ) ) # Print the RSpec to the enclosing page. portal.context.printRequestRSpec() Open this profile on CloudLab"""An example of using parameters to construct a profile with a variable number of nodes. Instructions: Wait for the profile instance to start, and then log in to one or more of the VMs via the ssh port(s) specified below. """ # Import the Portal object. import geni.portal as portal # Import the ProtoGENI library. import geni.rspec.pg as rspec # Describe the parameter(s) this profile script can accept. portal.context.defineParameter( "n", "Number of VMs", portal.ParameterType.INTEGER, 1 ) # Retrieve the values the user specifies during instantiation. params = portal.context.bindParameters() # Create a Request object to start building the RSpec. request = portal.context.makeRequestRSpec() # Check parameter validity. if params.n < 1 or params.n > 8: portal.context.reportError( portal.ParameterError( "You must choose at least 1 and no more than 8 VMs.", ["n"] ) ) # Abort execution if there are any errors, and report them. portal.context.verifyParameters() for i in range( params.n ): # Create a XenVM and add it to the RSpec. node = request.XenVM( "node" + str( i ) ) # Print the RSpec to the enclosing page. portal.context.printRequestRSpec()
Until now, all of the geni-lib scripts have described profiles which could also have been generated with the Jacks GUI, or even by writing a raw XML RSpec directly. However, geni-lib profiles offer an important feature unavailable by the other methods: the ability to describe not a static request, but a request “template” which is dynamically constructed based on a user’s choices at the time the profile is instantiated. The mechanism for constructing such profiles relies on profile parameters; the geni-lib script describes the set of parameters it will accept, and then retrieves the corresponding values at instantiation time and is free to respond by constructing arbitrarily different resource requests based on that input.
The profile above accepts exactly one parameter—
| Simple integer | |
| Arbitrary (uninterpreted) string | |
| True or False | |
| URN to a disk image | |
| URN of a GENI Aggregate Manager | |
| String specifying a type of node | |
| Floating-point number specifying bandwidth in kbps | |
| Floating-point number specifying delay in ms | |
| Integer used for memory or disk size (e.g., MB, GB, etc.) |
The last field is the default value of the parameter, and is required: not only must the field itself contain a valid value, but the set of all parameters must be valid when each of them assumes the default value. (This is partly so that the portal can construct a default topology for the profile without any manual intervention, and partly so that unprivileged users, who may lack permission to supply their own values, might still be able to instantiate the profile.)
After all parameters have been defined, the profile script may retrieve the runtime values with the bindParameters() method. This will return a Python class instance with one attribute for each parameter (with the name supplied during the appropriate defineParameter() call). In the example, the instance was assigned to params, and therefore the only parameter (which was called "n") is accessible as params.n.
Of course, it may be possible for the user to specify nonsensical values for a parameter, or perhaps give a set of parameters whose combination is invalid. A profile should detect error cases like these, and respond by constructing a portal.ParameterError object, which can be passed to the portal context’s reportError() method to abort generation of the RSpec. By default, reportError() does not abort the script immediately unless you pass it a specific argument; see its documentation for more detail. If you prefer to allow all errors and warnings to be generated, and to only abort the script at that point, and prior to creating RSpec resources, you can manually call verifyParameters() after completing parameter value checks. verifyParameters() is also automatically invoked prior to printing the request RSpec, so if your script can function despite malformed parameter values, you do not need to manually call verifyParameters().
8.9 Add storage to a node
Adding extra temporary diskspace to a node using a node-local dataset,
Creating a reusable dataset from local disk space using an image-backed dataset,
Adding network-attached storage to a node using a remote dataset,
Safely read-write sharing a remote dataset on multiple nodes via NFS,
Using read-write clones of a remote dataset on multiple nodes
8.10 Debugging geni-lib profile scripts
It is not necessary to instantiate the profile via the portal web interface
to test it. Properly written profile scripts should work perfectly well
independent of the normal portal—
python geni-lib-parameters.py
will produce an RSpec containing three nodes (the default value for n). It is also possible to override the defaults on the command line by giving the parameter name as an option, followed by the desired value:
python geni-lib-parameters.py –n 4
The option –help will list the available parameters and their descriptions.